Tn-Seq
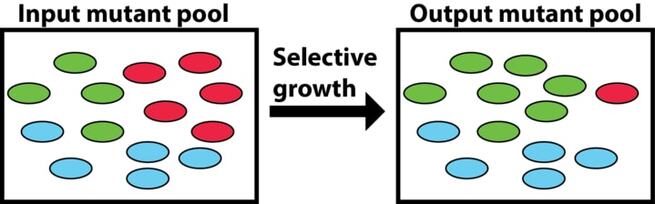
Transposon insertion site sequencing (Tn-seq) is a next-generation sequencing technique that allows simultaneous assessment of the relative frequencies of tens of thousands of individual transposon mutants following growth in specific environments. In addition, our lab has developed a Monte-Carlo based analysis of Tn-seq data that allows for the identification of essential genes.(Turner et al. 2015; Ibberson et al. 2017)
To successfully colonize and proliferate within a host organism, pathogens must overcome a variety of restrictions imposed by the host environment. Pathogens have evolved mechanisms to respond to environmental changes (e.g. acid or osmotic shock), acquire multiple carbon sources, and resist antimicrobial factors (e.g. produced endogenously by the host or supplied through antibiotic treatment). Identifying the genes required by pathogens during infection is central to understanding the biological processes controlling disease. We are interested in applying Tn-Seq to learn about mechanisms that enable the model pathogens Pseudomonas aeruginosa (Pa), Staphylococcus aureus (Sa) and Aggregatibacter actinomycetemcomitans (Aa) to survive these stresses.
Pa and Sa are infamous pathogens of the cystic fibrosis lung and chronic wounds, and Aa is implicated as a causative agent of periodontitis. These infections are frequently polymicrobial, and we have previously identified interspecies interactions that may worsen infection during co-colonization with Pa and Sa and with Aa and Streptococcus gordonii. We are using Tn-seq to develop genetic interaction networks and to identify essential genes of these bacteria during mono- and polymicrobial infection.
Pa and Sa are infamous pathogens of the cystic fibrosis lung and chronic wounds, and Aa is implicated as a causative agent of periodontitis. These infections are frequently polymicrobial, and we have previously identified interspecies interactions that may worsen infection during co-colonization with Pa and Sa and with Aa and Streptococcus gordonii. We are using Tn-seq to develop genetic interaction networks and to identify essential genes of these bacteria during mono- and polymicrobial infection.